Untying molecular knots: Making molecular simulations more efficient with LongBondEliminator
During lockdown, two things were on everyone’s mind: When is the vaccine going to come out, and while I’m waiting, what can I do to occupy my time? Some got into baking, others knitting. Researchers from the Vermaas lab at the MSU-DOE Plant Research Laboratory (PRL) combined these two questions and tried their hand at the puzzle that is vaccine molecular simulation. This had its challenges and led the researchers to create a more efficient tool to solve the problem of ring piercings in molecular simulations.
Molecular simulations use the power of computing to create a computational microscope which can observe biological structures on the atomic level. Researchers take structures they have observed in the lab and create computer models, which can involve fitting thousands of atoms together in just the right spots. It’s like working on a puzzle: With each new piece put into place, the overall picture of what the system is and what it can do becomes clearer.
“We have a general idea where the puzzle pieces, which means our molecular components, need to go,” said Martin Kulke, postdoc in the Vermaas lab, describing how these simulations are created. “But we use computer tools to automatically place the pieces, which leads to components overlapping. In the overlap of these puzzle pieces, we get the problem of these ring piercings.”
LongBondEliminator
By Josh Vermaas
Josh Vermaas, assistant professor at the PRL and the Department of Biochemistry & Molecular Biology and the primary investigator on this paper, was working on developing structural models for lignin, a polymer that is important in the plant cell wall. When creating lignin models that he could simulate, he was running into a common problem: In these simulations, the computer didn’t know what to do when one component overlapped with another and pierced the ring of the first, creating a model that would not be possible in reality.
When running into this issue, Vermaas and computational researchers like him would need to go through and fix all the errors by hand. This is a time consuming and tedious process.
“If we wanted to make good atomic models for these kinds of systems, there was a need to treat these problems in an automated way,” Vermaas said.
LongBondEliminator (LBE) is the tool Vermaas created to help fix these ring piercings with minimal user input.
“That started with lignin-specific code put into LigninBuilder, but LBE is generalizable to other systems with rings,” Vermaas said.
Improving on the tool
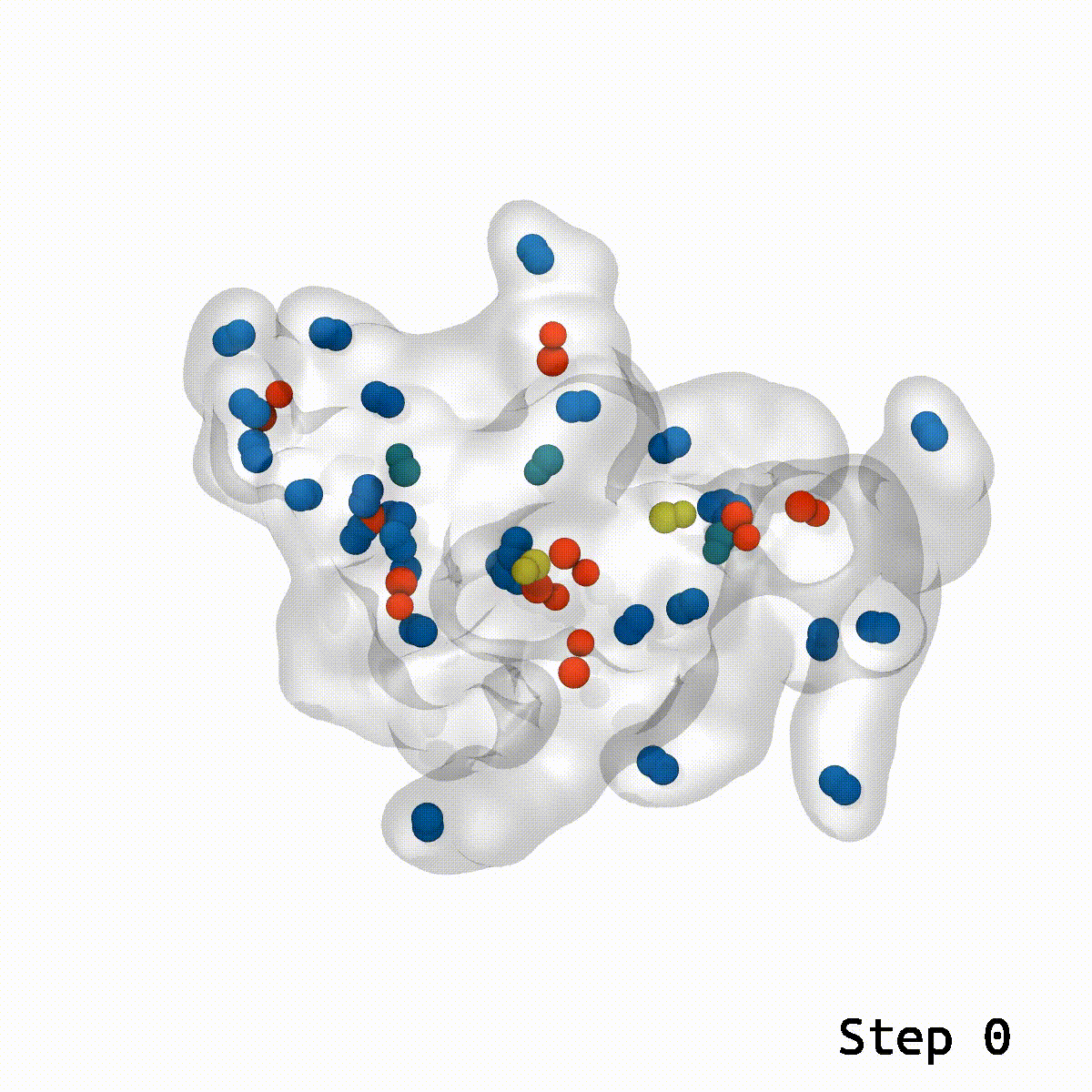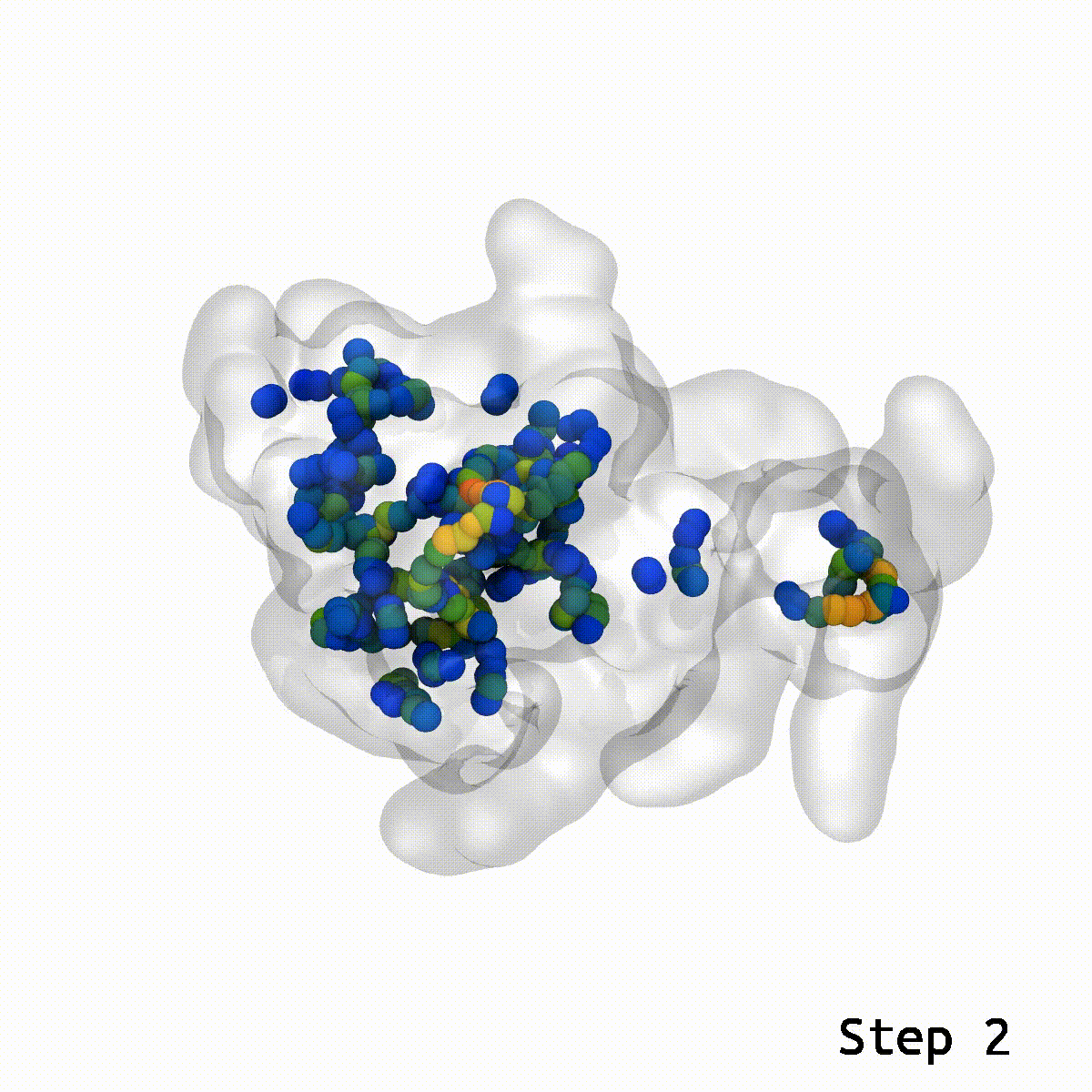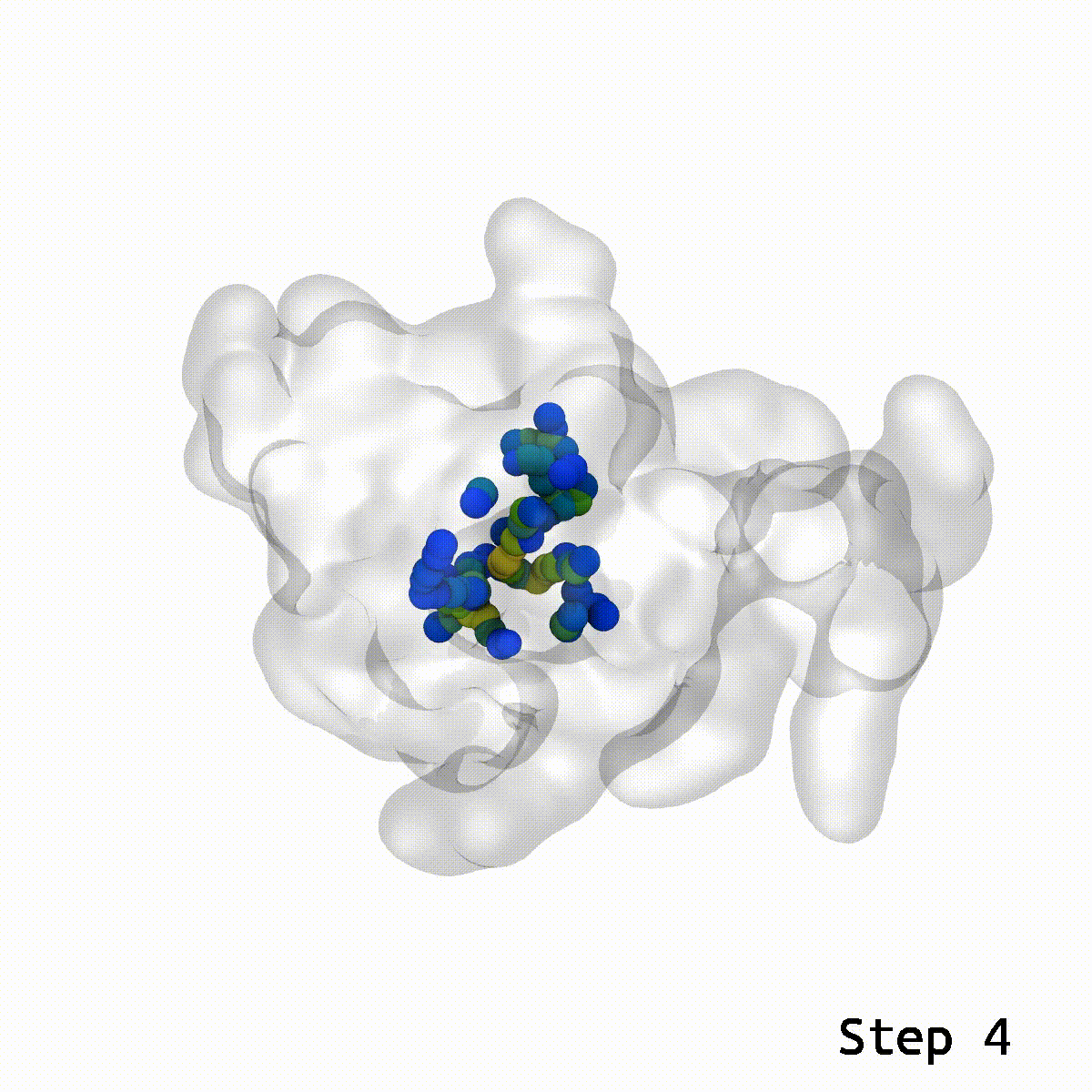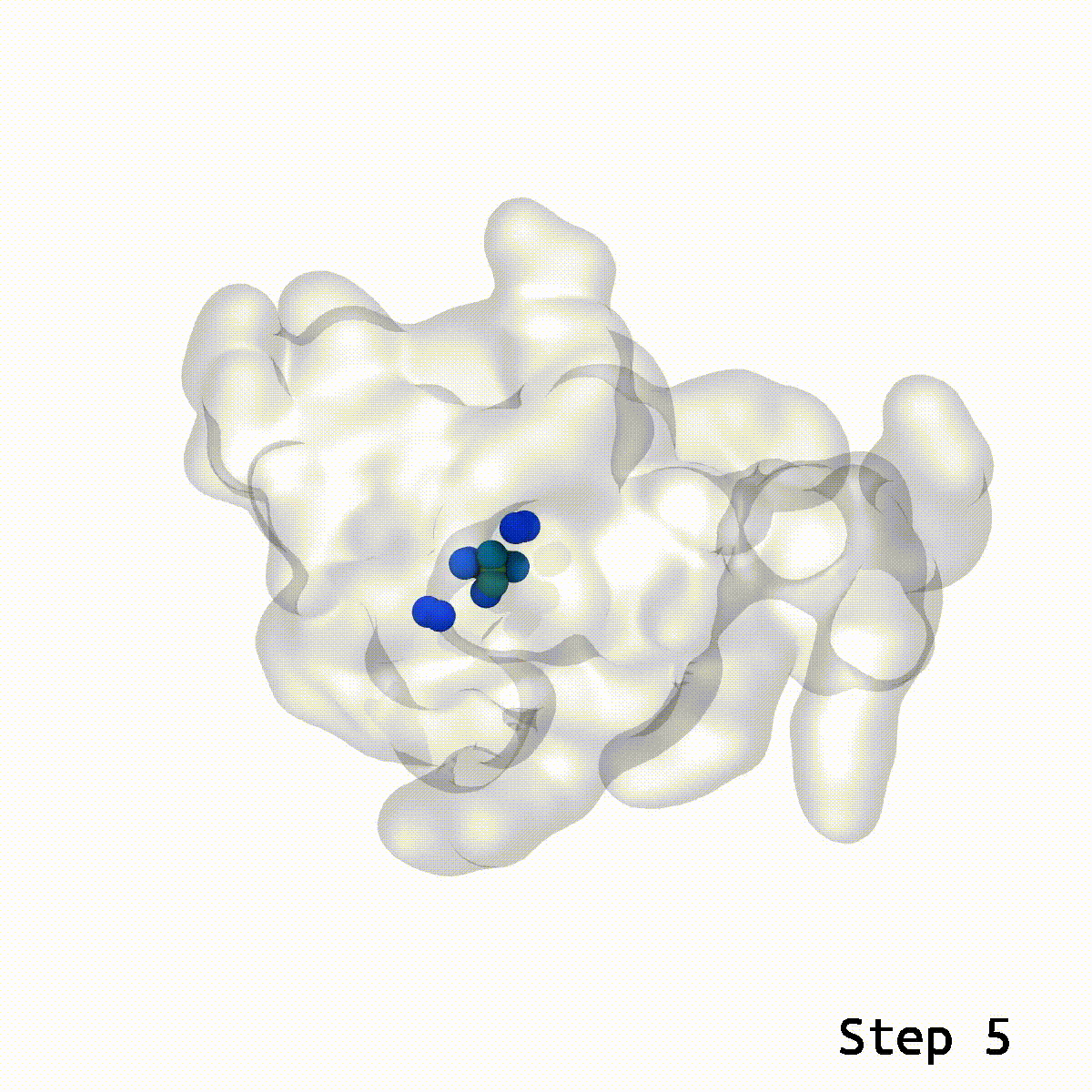
Animation rendered by Josh Vermaas
The three researchers were independently working on projects involving computational simulations, each with thousands of atoms that needed to be fit together. Kulke was simulating the glycosylated protein aggrecan, which is essential for the function of cartilage, and its degradation can result in diseases like arthritis. Postdoc Daipayan Sarkar was simulating the AstraZeneca COVID-19 vaccine with his collaborators, Abhishek Singharoy and Chun Kit Chan from Arizona State University and Alexander Baker at Accession Therapeutics, in collaboration with AstraZeneca to determine the structure of the ChAdOx1 COVID-19 viral vector (PDB: 7RD1).
When the researchers came together at the Vermaas lab, they soon realized they all had the same issues when it came to ring piercings. They worked to improve the LBE tool and make it more widely applicable in molecular simulations.
It was from the paper on the vaccine that the researchers in the Vermaas lab were invited by Alex Baker to write a paper for a special edition of Biomolecules, detailing their methods with LBE so that other researchers might utilize that tool efficiently.
“I would say that in any large system with many components that must come together, this tool can be important,” Kulke said.

Image 1 by Kara Headley; Images 2 and 3, courtesy photos
Sarkar added: “One of our goals at the PRL is to see the interaction between photosystem II and phycobilisome in cyanobacteria. These two systems need to be put together; the phycobilisome needs to be stacked on top of photosystem II. We don’t have enough information at the atomic level to do this completely, but we can create a starting model which we can keep building on and derive biochemical and biophysical information from. When we do this, we expect to have problems with ring piercings.”
This is where the LBE comes into play. With a reasonable guess for the relative orientation between two structures, LBE makes it simple to eliminate these show-stopping artifacts.
“The real key is that we want to be spending our time doing cool science, setting up accurate models for the types of questions of interest to the PRL,” Vermaas said. “Creating bespoke solutions for individual systems was just much too inefficient."
This work was supported by the Division of Chemical Sciences, Geosciences and Biosciences, Office of Basic Energy Sciences of the U.S. Department of Energy Grant DE-FG02-91ER20021.
By Kara Headley